7.5 Example
Returning to the Lung function data set from PMA6, lets analyze the relationship between height and FEV for fathers in this data set.
ggplot(fev, aes(y=FFEV1, x=FHEIGHT)) + geom_point() +
xlab("Height") + ylab("FEV1") +
ggtitle("Scatter Diagram with Regression (blue) and Lowess (red) Lines
of FEV1 Versus Height for Fathers.") +
geom_smooth(method="lm", se=FALSE, col="blue") +
geom_smooth(se=FALSE, col="red") 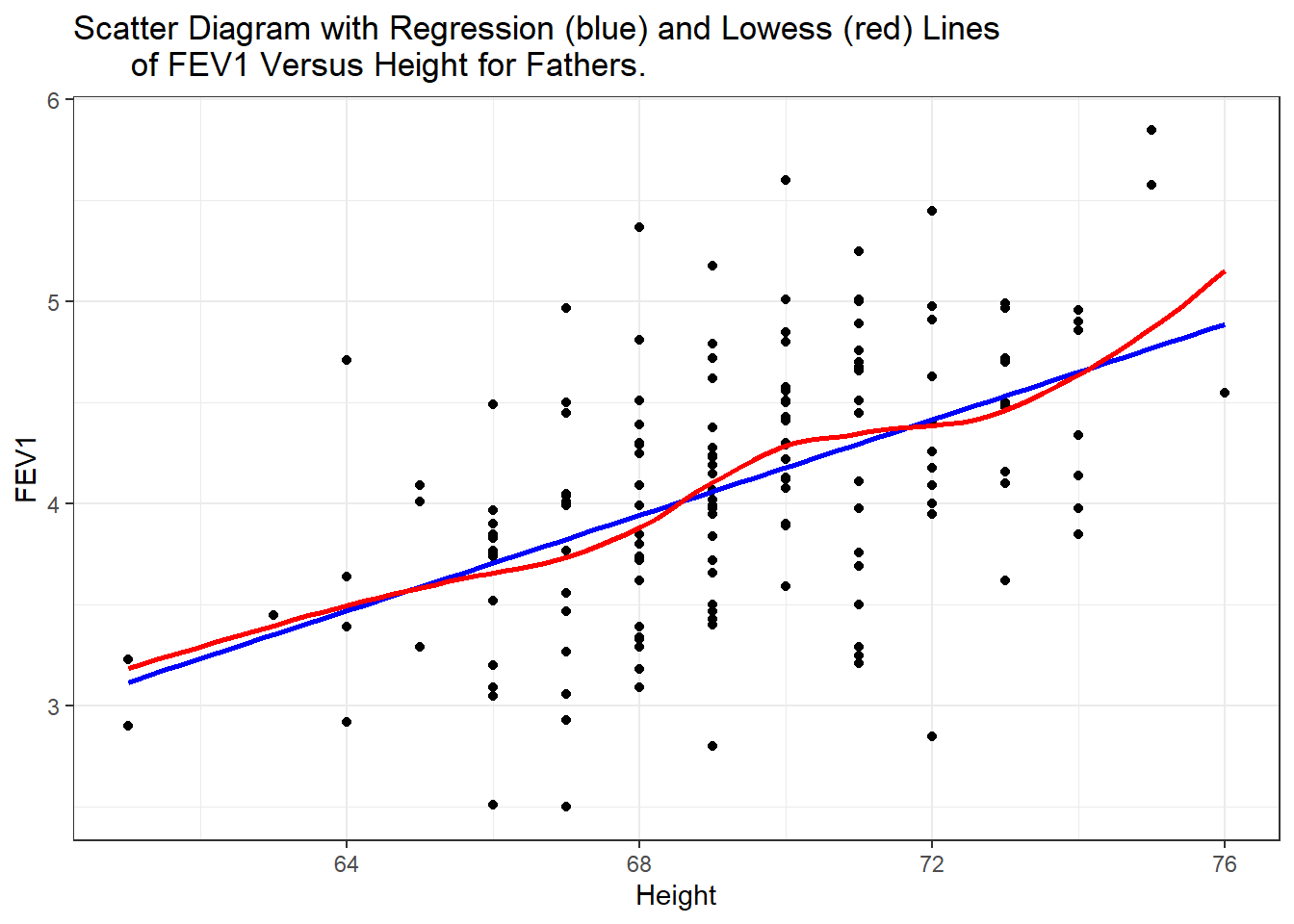
There does appear to be a tendency for taller men to have higher FEV1. The trend is linear, the red lowess trend line follows the blue linear fit line quite well.
Let’s fit a linear model and report the regression parameter estimates.
| term | estimate | std.error | statistic | p.value |
|---|---|---|---|---|
| (Intercept) | -4.087 | 1.152 | -3.548 | 0.001 |
| FHEIGHT | 0.118 | 0.017 | 7.106 | 0.000 |
The least squares equation is \(Y = -4.09 + 0.118X\). We can calculate the confidence interval for that estimate using the confint function.
| 2.5 % | 97.5 % | |
|---|---|---|
| (Intercept) | -6.363 | -1.810 |
| FHEIGHT | 0.085 | 0.151 |
For ever inch taller a father is, his FEV1 measurement significantly increases by .12 (95%CI: .09, .15, p<.0001).